You are here: Vanderbilt Biostatistics Wiki>Main Web>Projects>MicroArrayMassSpec>GeneralWfccmDesign>WfccmClassDescriptions>WfccmAlgorithmHu (07 May 2009, WillGray)Edit Attach
Hu Algorithm
function 1 : log-likelihood to estimate logmu -
f(logmu)=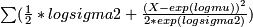
- logsigma2 = beta0 + beta1 * logmu
function 2 : log-likelihood to estimate beta -
f(beta)=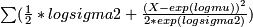
- logsigma2 = beta[1] + beta[2] * logmu
Data Preprocessing
- seperate the original data set to two matrix.
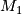
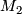
- calculate rchisq

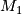
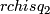
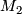
- calculate variance
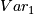

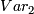
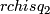
- calculate 1% quantile
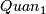
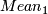
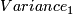
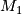 ,
, 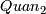
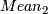
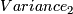
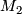 without NAN and negative values
without NAN and negative values
n is sample of population, x is value corresponding to each sample
(a)mean=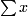 / n
/ n
(b)variance=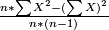
(c)1% quantile= value at (pos-1) + diff_v*diff_p
position = 1 + (n-1)*0.01
pos = integer part of position
diff_p = position - pos
diff_v = ceiling of value at (position - 1 ) - value at (pos-1)
- replace NA and negative values in
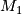 with
with 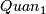 ,
, 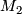 with
with 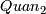
Estimate MLE
- calculate coefficients and residuals based on log values of
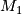 and
and 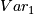 ,
, 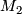 and
and 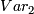
x are values in M, y are values in Quan
coefficients:
(a) intercept of coefficient =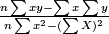
(b) slope of coefficient = y - intercept * x
residuals= y - (intercept + slope * x)
-
xd2hat= variance of residuals - Var
- calculate initial values
xinitial= estimate value ofnlm - lkdsumsloop loop 3 times to calculate
lkdinitial
if lkdinf-lkdsumsloop[3] >= 1
then lkdsumsloop== lkdsumsloop[-1]
(a) calaulateminandestimateof xinitial
(b) lkdinitial =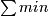
- calculate
lkdinf
lkdinf = lkdsumsloop[1]+![\begin{displaymath} \frac{1}{(1-\frac{lkdsumsloop[3]-lkdsumsloop[2]}{lkdsumsloop[2]-lkdsumsloop[1]})*(lkdsumsloop[2]-lkdsumsloop[1])} \end{displaymath}](/wiki/pub/Main/WfccmAlgorithmHu/latexbe23cfd4108995a329066fe6310cac5e.png)
- calcualte mle estimate of mean expression, mle estimate of variance of each gene
(a)xbeta.mle= xinitial[(m+1):(m+2)]
(b)xbar.mle= exponent of xinitial
(c)xs2.mle= exponent of xbeta.mle * xbar.mle ^ xbeta.mle
calculate Hu score
score = |Remarks
- Details of nlm calculation have been skipped.
Edit | Attach | Print version | History: r13 < r12 < r11 < r10 | Backlinks | View wiki text | Edit wiki text | More topic actions
Topic revision: r13 - 07 May 2009, WillGray
 Copyright &© 2013-2022 by the contributing authors. All material on this collaboration platform is the property of the contributing authors.
Copyright &© 2013-2022 by the contributing authors. All material on this collaboration platform is the property of the contributing authors. Ideas, requests, problems regarding Vanderbilt Biostatistics Wiki? Send feedback
