You are here: Vanderbilt Biostatistics Wiki>Main Web>Education>HandoutsBioRes>ClinStat>ManuscriptChecklist>BootstrapMeansSoftware (09 Mar 2007, FrankHarrell)Edit Attach
Software and Example Code for Bootstrap Confidence Intervals for Means and Differences in Means
The following uses the simple nonparametric bootstrap percentile approach inR. To obtain 0.95 confidence intervals for a single mean use
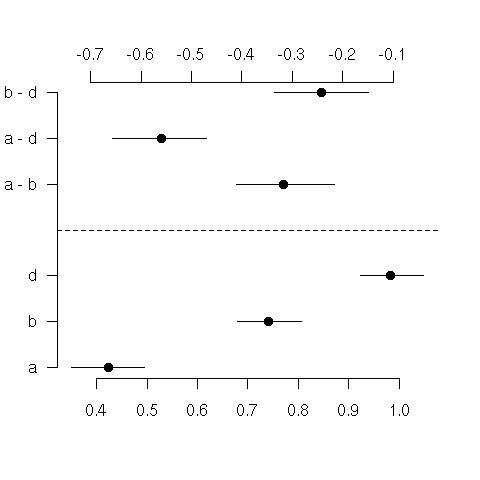
Bootstrap confidence intervals for means and differences in means
| I | Attachment | Action | Size | Date | Who | Comment |
|---|---|---|---|---|---|---|
| |
bootdiffs.png | manage | 2 K | 09 Mar 2007 - 11:12 | FrankHarrell | Bootstrap confidence intervals for means and differences in means |
Edit | Attach | Print version | History: r1 | Backlinks | View wiki text | Edit wiki text | More topic actions
Topic revision: r1 - 09 Mar 2007, FrankHarrell
 Copyright &© 2013-2022 by the contributing authors. All material on this collaboration platform is the property of the contributing authors.
Copyright &© 2013-2022 by the contributing authors. All material on this collaboration platform is the property of the contributing authors. Ideas, requests, problems regarding Vanderbilt Biostatistics Wiki? Send feedback


